This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison, Spring 2012.
Phylogenic trees from DNA sequence
What is a phylogeny?
A phylogeny is a depiction of changing organismal lineages over time, where branch points represent a common ancestor of the two diverging species. Using the degree of similarity between homologous gene sequences allows us to estimate how related species are to each other and to construct the phylogenetic trees seen below.
Phylogeny.fr Tree
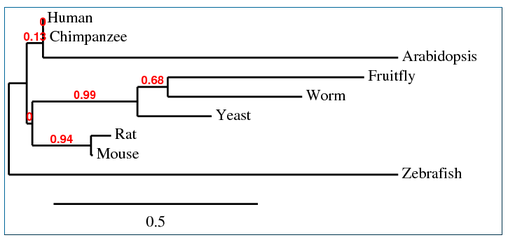
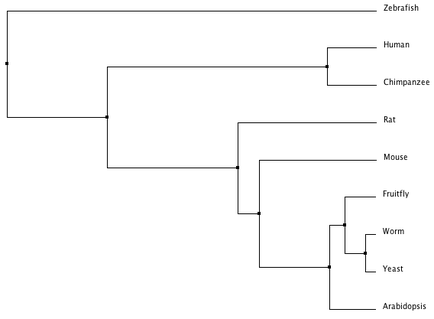
This tree from the alignment of Homo sapiens SOD1 gene with homologs in P. troglodytes, M. musculus,
R. norvegicus, D. rerio, D. melanogaster, A. thaliana, C. elegans and S. cerevisiae was generated by Phylogeny.fr using default settings in "one click" mode [1,2,3,4,5,6,7].
R. norvegicus, D. rerio, D. melanogaster, A. thaliana, C. elegans and S. cerevisiae was generated by Phylogeny.fr using default settings in "one click" mode [1,2,3,4,5,6,7].
T-Coffee Phylogeny
The following tree was created by aligning DNA sequences in T-Coffee using default settings [8,9]. The tree was calculated using average distance based on percent identity.
MUSCLE Phylogeny
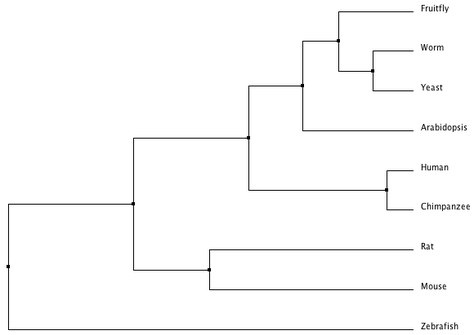
The following tree was created by aligning DNA sequences in MUSCLE using default settings [3]. It was calculated using average distance based on percent identity.
Analysis
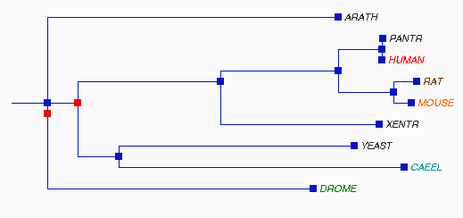
Phylogeny.fr, T-Coffee, and MUSCLE all created different trees. The phylogeny constructed at Phylogeny.fr unexpectedly put Arabidopsis SOD1 as the closest homolog related to humans and chimpanzees. The T-Coffee and MUSCLE trees are more similar to each other and to the trees based on more accurate alignments using the protein sequences. The differences between the Phylogeny.fr and MUSCLE alignment are interesting because Phylogeny.fr uses MUSCLE alignments in its process. The T-Coffee and MUSCLE trees are also closest to the following tree from the TreeFam database for the SOD1 gene comparing the same organisms as above (with frogs instead of zebrafish) [10,11]. TreeFam trees are created from DNA sequences and then manually curated using online databases and journal articles to correct for errors, which should make them more reliable.
While Phylogeny.fr is the most user friendly of these systems, in that you paste your sequences and get a tree, I think the little bit of extra work in exploring T-Coffee or MUSCLE results to make a tree is worth it for a more accurate result. Searching the TreeFam database is very informative, but the extent of the trees can be overwhelming. Since they all use different algorithms, using multiple systems seems to be the most reliable approach.
- Dereeper A., Audic S., Claverie J.M., Blanc G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol Biol. 2010 Jan 12;10:8. PMID:20067610
- Dereeper A.*, Guignon V.*, Blanc G., Audic S., Buffet S., Chevenet F., Dufayard J.F., Guindon S., Lefort V., Lescot M., Claverie J.M., Gascuel O. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008 Jul 1;36(Web Server issue):W465-9. Epub 2008 Apr 19. PMID:18424797
- Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, Mar 19;32(5):1792-7. PMID:15034147
- Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol. 2000, Apr;17(4):540-52. PMID:10742046
- Guindon S., Gascuel O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 2003, Oct;52(5):696-704. PMID:14530136
- Anisimova M., Gascuel O. Approximate likelihood ratio test for branchs: A fast, accurate and powerful alternative. Syst Biol. 2006, Aug;55(4):539-52. PMID:16785212
- Chevenet F., Brun C., Banuls AL., Jacq B., Chisten R. TreeDyn: towards dynamic graphics and annotations for analyses of trees. BMC Bioinformatics. 2006, Oct 10;7:439. PMID:17032440
- Di Tommaso P, Moretti S, Xenarios I, Orobitg M, Montanyola A, Chang JM, Taly JF, Notredame C. (2011). T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 39:W13-7. PMID:21558174
- Notredame, C., Higgins, D., Heringa, J. (2000). T-Coffee: A novel method for fast and accurate multiple sequence alignments. JMB 302: 205-217. PMID:10964570
- Li, H. et al. (2006). TreeFam: a curated database of phylogenic trees of animal gene families. Nucleic acid res. 34: D572-80. PMID:16381935
- Ruan, J. et al. (2008). TreeFam: 2008 Update. Nuclei acid res. 36: D735-40. PMID:18056084