This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison, Spring 2012.
SOD1 Mutant and RNAi Phenotypes
Mutant phenotypes in model organisms
Organism databases are a quick way to look up information on mutant phenotypes in difference organisms.
Since model organisms are used to study ALS in humans, it is important to understand the similarities and differences in SOD1 mutant and RNAi phenotypes in humans versus model organisms. Below are mutant and RNAi phenotypes from model organisms with links to the databases at the right.
|
Mouse mutant phenotypes from MGI [1]:
SOD1 homozygous mutants exhibit increased motor neuron loss after neuron injury. Females have irregular and small litters, and for some alleles exhibit immature ovarian follicles. Zebrafish mutant phenotypes from ZFIN [2]: SOD1 mutants have disrupted sensory perception of touch and in concert with other mutations can lead to a decrease in motor axon length. Fruit fly mutant phenotypes from Flybase [3]: SOD mutants have a reduced lifespan and show defects in oxidative stress response. Worm mutant phenotypes from Wormbase [4]: SOD-1 mutants show extended lifespan, variant protein folding response and variant protein metabolism. Yeast mutant phenotypes from SGD [5]: SOD1p mutants show phenotypes including decreased lifespan, decreased oxidative stress response, decreased alkaline pH response and decreased sporulation. |
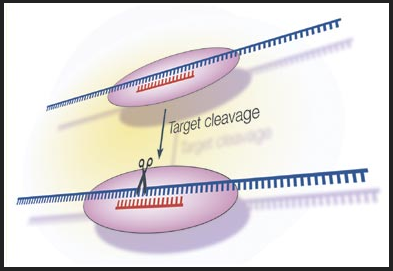
What is RNA interference (RNAi)?
RNAi is a technique used in many organisms to knockdown gene function through the degradation of targeted RNAs. Using RNAi can produce phenotypes similar to mutant phenotypes.
Below are RNAi phenotypes in model organisms where available:
Fruit Fly RNAi phenotypes from Genome RNAi via Flybase:
Using sod RNAi, researchers saw defects in lipid storage and bristle morphology as well as decreased viability of flies.
Worm RNAi phenotypes from Wormbank:
Using sod-1 RNAi, researchers saw defects in organism development, body wall muscle myosin organization, and egg laying, as well as multivulva phenotypes and decreased lifespan.
Fruit Fly RNAi phenotypes from Genome RNAi via Flybase:
Using sod RNAi, researchers saw defects in lipid storage and bristle morphology as well as decreased viability of flies.
Worm RNAi phenotypes from Wormbank:
Using sod-1 RNAi, researchers saw defects in organism development, body wall muscle myosin organization, and egg laying, as well as multivulva phenotypes and decreased lifespan.
Analysis
Mice phenotypes are fairly similar to human phenotypes, making mice a great model organism for studying ALS. In mice, mutants can be created that match known human mutations because of the similar protein homology. Because these directed mutations are so effective, RNAi experiments are less common in mice, except for when testing potential uses for sod1 RNAi in ALS treatment.
In the other model organisms listed, mutants in sod1 homologs show defects in handling oxidative stress and decreased lifespan, so ALS can also be studied in these organisms, especially looking at the mechanisms of how mutant SOD1 disrupts cellular function.
Model organism databases are useful tools for finding a wide range of information about your gene of interest in different organisms, from gene sequences, to mutant phenotypes, to journal articles referencing the gene.
In the other model organisms listed, mutants in sod1 homologs show defects in handling oxidative stress and decreased lifespan, so ALS can also be studied in these organisms, especially looking at the mechanisms of how mutant SOD1 disrupts cellular function.
Model organism databases are useful tools for finding a wide range of information about your gene of interest in different organisms, from gene sequences, to mutant phenotypes, to journal articles referencing the gene.
1. Blake JA, Bult CJ, Kadin JA, Richardson JE, Eppig JT and the Mouse Genome Database Group. 2011. The Mouse Genome Database (MGD): premier model organism resource for mammalian genomics and genetics. Nucleic Acids Res 39(suppl 1): D842-D848. PMID:21051359
2. Bradford, Y., Conlin, T., Dunn, N., Fashena, D., Frazer, K., Howe, D.G., Knight, J., Mani, P., Martin, R., Moxon, S.A., Paddock, H., Pich, C., Ramachandran, S., Ruef, B.J., Ruzicka, L., Bauer Schaper, H., Schaper, K., Shao, X., Singer, A., Sprague, J., Sprunger, B., Van Slyke, C., and Westerfield, M. (2011) ZFIN: enhancements and updates to the zebrafish model organism database. Nucleic Acids Res.. 39(suppl 1):D822-D829.
PMID:21036866
3. Peter McQuilton, Susan E. St. Pierre, Jim Thurmond, and the FlyBase Consortium (2012).
FlyBase 101 – the basics of navigating FlyBase. Nucleic Acids Res. 40(Database issue):D706-14.
PMID:22127867
4. Lincoln D. Stein, Paul Sternberg, Richard Durbin, Jean Thierry-Mieg, and John Spieth (2001). WormBase: network access to the genome and biology of Caenorhabditis elegans. Nucleic Acids Research 29:82-86.
PMID:11125056
5. Cherry JM, Hong EL, Amundsen C, Balakrishnan R, Binkley G, Chan ET, Christie KR, Costanzo MC, Dwight SS, Engel SR, Fisk DG, Hirschman JE, Hitz BC, Karra K, Krieger CJ, Miyasato SR, Nash RS, Park J, Skrzypek MS, Simison M, Weng S, Wong ED (2012) Saccharomyces Genome Database: the genomics resource of budding yeast. Nucleic Acids Res. Jan;40(Database issue):D700-5. PMID:22110037
2. Bradford, Y., Conlin, T., Dunn, N., Fashena, D., Frazer, K., Howe, D.G., Knight, J., Mani, P., Martin, R., Moxon, S.A., Paddock, H., Pich, C., Ramachandran, S., Ruef, B.J., Ruzicka, L., Bauer Schaper, H., Schaper, K., Shao, X., Singer, A., Sprague, J., Sprunger, B., Van Slyke, C., and Westerfield, M. (2011) ZFIN: enhancements and updates to the zebrafish model organism database. Nucleic Acids Res.. 39(suppl 1):D822-D829.
PMID:21036866
3. Peter McQuilton, Susan E. St. Pierre, Jim Thurmond, and the FlyBase Consortium (2012).
FlyBase 101 – the basics of navigating FlyBase. Nucleic Acids Res. 40(Database issue):D706-14.
PMID:22127867
4. Lincoln D. Stein, Paul Sternberg, Richard Durbin, Jean Thierry-Mieg, and John Spieth (2001). WormBase: network access to the genome and biology of Caenorhabditis elegans. Nucleic Acids Research 29:82-86.
PMID:11125056
5. Cherry JM, Hong EL, Amundsen C, Balakrishnan R, Binkley G, Chan ET, Christie KR, Costanzo MC, Dwight SS, Engel SR, Fisk DG, Hirschman JE, Hitz BC, Karra K, Krieger CJ, Miyasato SR, Nash RS, Park J, Skrzypek MS, Simison M, Weng S, Wong ED (2012) Saccharomyces Genome Database: the genomics resource of budding yeast. Nucleic Acids Res. Jan;40(Database issue):D700-5. PMID:22110037