This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison, Spring 2012.
SOD1 Protein Homologs and Alignments
What is a homolog?
A homolog is a protein in one organism that is related to a protein in another organism by descent from a common ancestor. Therefore, homologs can help us to better understand similarities and differences between organisms on the sub-cellular level. A protein that is well conserved likely has an important and similar function in multiple organisms. Because of the redunancy of the genetic code, homologs are expected to be more similar when comparing amino acid sequences that make up proteins than when comparing the genomic DNA sequences.
Identifying homologs is a critical step in the study of human diseases, as the early research to understand and treat diseases like ALS is typically done using model organisms.
Identifying homologs is a critical step in the study of human diseases, as the early research to understand and treat diseases like ALS is typically done using model organisms.
Protein Sequences
The SOD1 protein is well conserved throughout most species. Below are the links to protein sequences of Human SOD1 homologs in selected model organisms:
|
Pan troglodytes (Chimpanzee)
Superoxide dismutase 1 (SOD1): NP_001009025.1 length: 154 aa Mus musculus (Mouse) Superoxide dismutase 1 (SOD1): NP_035564.1 length: 154 aa Rattus norvegicus (Rat) Superoxide dismutase 1 (SOD1): NP_058746.1 length: 154 aa Danio rerio (Zebrafish) Superoxide dismutase 1 (SOD1): NP_571369.1 length: 154 aa |
Drosophila melanogaster (Fruit Fly)
Superoxide dismutase (SOD): NP_476735.1 length: 153 aa Caenorhabditis elegans (Roundworm) Superoxide dismutase (SOD): NP_001021957.1 length: 158 aa Saccharomyces cerevisiae (Yeast) Superoxide dismutase 1 (SOD1P): NP_012638.1 length: 154 aa Arabidopsis thaliana (Mouse-ear Cress) CSD1: NP_172360.1 length: 152 aa |
ClustalW Alignment
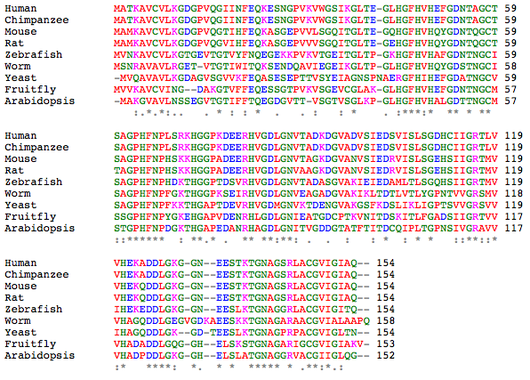
The following alignment of the above proteins was generated by ClustalW using default settings [1,2]. For EBI's explanation of the color scheme and consensus symbols, click here.
|
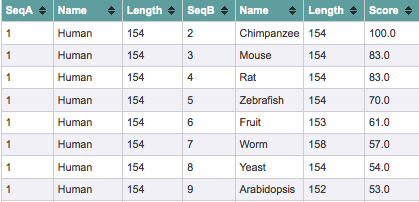
Below are the ClustalW scores for the alignment of human SOD1 to the selected homologs. Score is a measure of percent identity, or the number of identical residues between the two sequences being compared. Human and chimpanzee SOD1 are 100% identical while human SOD1 and its least similar homolog, Arabidopsis SOD1, are 53% identical. To view all of the scores of each homolog aligned to every other, click here.
|
T-Coffee Expresso Alignment
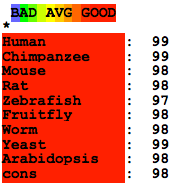
Below is a summary of the accuracy of the T-Coffee Expresso alignment of the homologous proteins. This algorithm takes into account structural information know about the proteins. Click here to download the alignment [3,4]. In this format, the colors and scores to the right denote the predicted accuracy of the alignment. Note the contrast between the accuracy of the protein alignment and the DNA sequence alignment.
Analysis
Aligning the protein sequences from these organisms shows great similarity between species - much more than is seen for the DNA sequences themselves. Human SOD1 is identical to chimpanzee SOD1 and least similar to Arabidopsis SOD1, sharing 53% identity. Since the proteins share so much similarity, the ClustalW alignment based on sequence and the T-Coffee Expresso alignment, taking known structure into consideration, are very similar and predicted to be quite accurate. The residues conserved throughout all of these organism include those involved in Zn and Cu binding and the disulfide bond.
1. Larkin M, Blackshields G, Brown N, Chenna R, McGettigan P, McWilliam H, Valentin F, Wallace I, Wilm A, Lopez R, Thompson J, Gibson T, Higgins D. (2007.) Clustal W and Clustal X Version 2.0. Bioinformatics, 23, 2947-2948. PMID:17846036
2. Thompson J, Higgins D, Gibson T. (1994.) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res., 22, 4673-4680. PMID:7984417
3. Di Tommaso P, Moretti S, Xenarios I, Orobitg M, Montanyola A, Chang JM, Taly JF, Notredame C. (2011.) T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 39:W13-7. PMID:21558174
4. Notredame, C., Higgins, D., Heringa, J. (2000.) T-Coffee: A novel method for fast and accurate multiple sequence alignments. JMB 302: 205-217. PMID:10964570
2. Thompson J, Higgins D, Gibson T. (1994.) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res., 22, 4673-4680. PMID:7984417
3. Di Tommaso P, Moretti S, Xenarios I, Orobitg M, Montanyola A, Chang JM, Taly JF, Notredame C. (2011.) T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 39:W13-7. PMID:21558174
4. Notredame, C., Higgins, D., Heringa, J. (2000.) T-Coffee: A novel method for fast and accurate multiple sequence alignments. JMB 302: 205-217. PMID:10964570