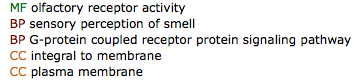This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison, Spring 2012.
SOD1 DNA Motifs
What is a DNA motif?
DNA sequence motifs are short, recurring DNA sequences that are presumed to have some biological function [1]. They can have many functions such as transcription factor binding sites at the DNA level or function in RNA splicing, just to name a couple. For more information about sequence motifs and how they are discovered and represented, check out this primer.
Conserved SOD1 DNA Sequence Motifs
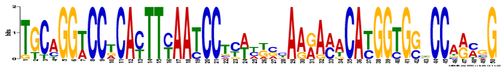
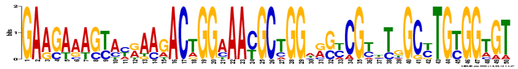
MEME identified 3 conserved DNA motifs in the full length human SOD1 DNA sequence and 8 of its homologs using default settings and searching for motifs that occur 0 or 1 time per sequence. (For SOD1 sequences, see gene homology.) In this representation, the height of each nucleotide in each position represents the frequency that it occurs in that position. A smaller E-value represents higher confidence [2].
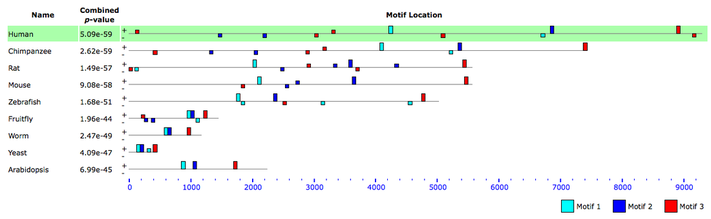
The following graphic displays the positions of these motifs in the sequence of each homolog. The blocks are proportional to the -log(p-value), so long blocks above the mid-line are the best matches. (Click to enlarge.)
Analysis
MEME allows for the discovery of similar DNA sequence motifs across multiple sequences, and provides links to multiple sources to analyze these motif. Since these motifs are conserved across organisms, they may serve some important function. Therefore, the sequences above were analyzed using GOMO: Gene Ontology for Motifs in order to connect the motifs to potential gene ontology terms. Below are the GO terms listed for motif 1 above when linked directly from the MEME results page using the multiple species, Homo sapiens database [3]. GOMO failed to identify significant GO terms for motifs 2 and 3 above.
While these GO terms do not correlate directly with GO terms associated with SOD1 (see gene ontology), they may provide insight into abnormal processes of mutant SOD1 and its toxic gain-of-function properties.
1. D’haeseleer P. (2006.) What are DNA sequence motifs? Nature Biotech 24: 423-425. PMID:16601727
2. Bailey T and Elkan C. (1994.) Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California. PMID:7584402
3. Buske F, Boden M, Bauer D and Bailey T. (2010.) Assigning roles to DNA regulatory motifs using comparative genomics. Bioinformatics 26(7): 860-866. PMID:20147307
2. Bailey T and Elkan C. (1994.) Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California. PMID:7584402
3. Buske F, Boden M, Bauer D and Bailey T. (2010.) Assigning roles to DNA regulatory motifs using comparative genomics. Bioinformatics 26(7): 860-866. PMID:20147307